Entering edit mode
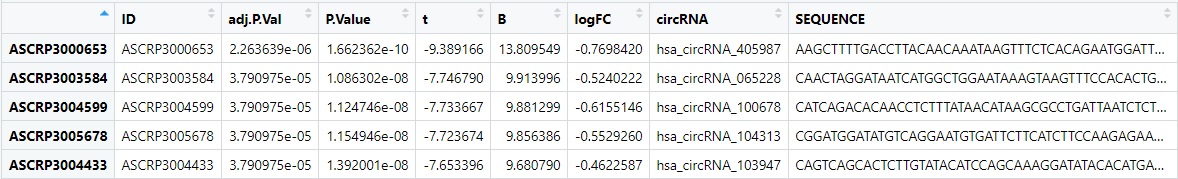
Hi! I'm wondering how to retrieve, from a given circRNA ( e.g. hsa_circRNA_405987), the genomic position in human genome 19 coordinates ( e.g. 121765368) using R.
Do you have any ideas about it?
Should I do a preliminary step from circRNA to transcript and then from transcript to gene SYMBOL? Something like that?
Many thanks!