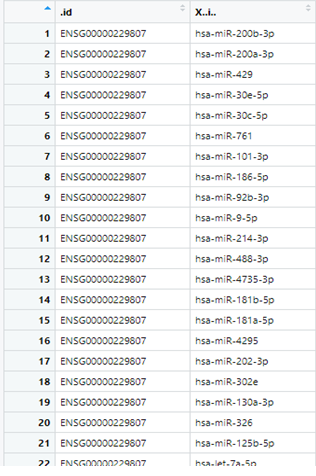
Hi, I have a question regarding converting the data frame to a nested list in R. I have a data frame as shown in the screenshot below wherein column 1 depicts the Ensembl IDs and column 2 are the miRNAs.
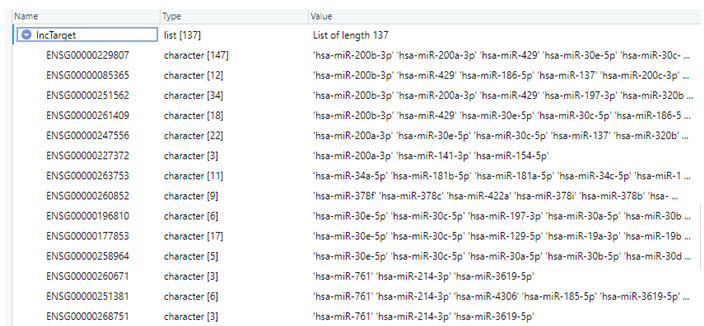
Now, I want to convert this data frame to a nested list such that the redundancy in column 1 (i.e., Ensembl IDs) is removed and the list looks something like this as follows:
It would be really great if you can please provide any suggestions or recommendations to accomplish this in R. Thanks very much!


Quite simply, you can use the nest function. This will create a column that contains a tibble for the data associated with each ID.
Sometimes it's handy to have that list column in a different form, such as an array or a JSON string (e.g. including in a database). Alternatively, you may want the data in a list as opposed to a tibble.
Here are some examples:
Created on 2021-12-12 by the [reprex package](https://reprex.tidyverse.org) (v2.0.1)
Hi Ariel, thanks very much for responding. I really appreciate it. Have a good one!