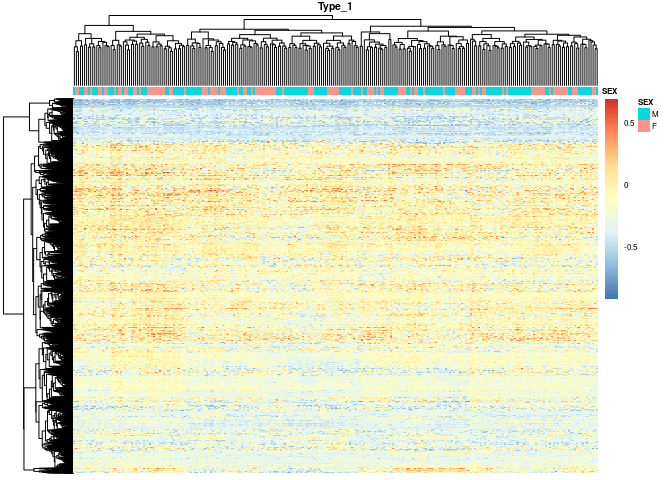
I am running GSVA on normalized RNA seq data. Is it acceptable to get all negative enrichment scores for a pathway? The heatmap below shows GSVA scores with pathways in rows and samples in columns. We note that some rows are all blue (i.e., enrichment scores for all samples for that pathway are negative). For those blue pathways, distributions of enrichment scores are unimodal and approx. symmetric with mean < 0. Can this happen? Or is a pathway supposed to have both positive and negative enrichment scores? In the paper, it appears that kernels are calculated, accounting for other samples' expression level. I am not sure if the results can be trusted. Please help! Thanks!
hi, if the question is about what a GSVA negative score means, this has been answered before, please check this post. If the question is why some pathways get negative GSVA scores in all your samples, you'll find the answer by looking into the genes that form those pathways. In principle, those genes should be ranking low in their expression across all the samples. One possibility is that they are non-expressed genes in all those samples, with consequently very low values of expression. If you have not done it, I strongly recommend you to filter out lowly-expressed genes before using GSVA, just as you normally do before doing a differential expression analysis.
cheers,
robert.
Use of this site constitutes acceptance of our User Agreement and Privacy Policy.