Dear Sir/Madam,
my goal is to Identify of the peptide differentially expressed between pre and post SARS-CoV-2 infection samples.
My count matrix is compose of 100 samples (paired, 50 samples before infection, and the same 50 samples after infection) with 3 million measured peptide (CPM). The data distribution is poisson-like distribution.
I have several question with regard to use of LIMMA package for the my purpose:
1) In general, Is the limma an optimal method to use for this kind of data and my purpose?
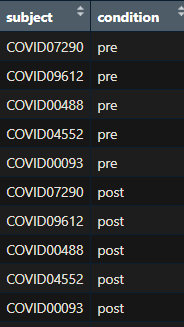
2) If this is the case what should be my design matrix? is this design correct? ( The screenshot of my sample information is added)
TA = factor(sample_info$subject) SA = factor(sample_info$condition, levels = c("pre","post")) design <- model.matrix(~TA+SA)
3) Should be the order of samples in sample information data frame the same as the order of samples in count matrix (I mean the column order in the count matrix)?
4) My code is like the following:
dge <- DGEList(counts=bigdf_t) keep <- filterByExpr(dge, design) dge <- dge[keep,,keep.lib.sizes=FALSE]
dge <- calcNormFactors(dge) v <- voom(dge, design, plot=FALSE) fit <- lmFit(v, design) fit <- eBayes(fit) topTable(fit, coef=ncol(design))
What should be the parameter in coef in topTable? should it be the last column in design matrix which basically shows the pre and post in condition?
5) What the final result shows me is the top 10 peptide which potentially differentially expressed with different p-value but the same adjusted p-value (0.9999996). the method for adjusting p-value is BH. I want to know how I can interpret this result? Does it mean no peptide is really differentially expressed or there is sth wrong with my code/data?
I would appreciate all of your kind help in advance.