Hi everyone, I currently have a dataset of 6 samples. The experimental design is 3 conditions, with 2 samples in each condition.
Therefore, my coldata table looks like this:
Samp_ID Condition
S1 Cond_1
S2 Cond_1
S3 Cond_2
S4 Cond_2
S5 Cond_3
S6 Cond_3
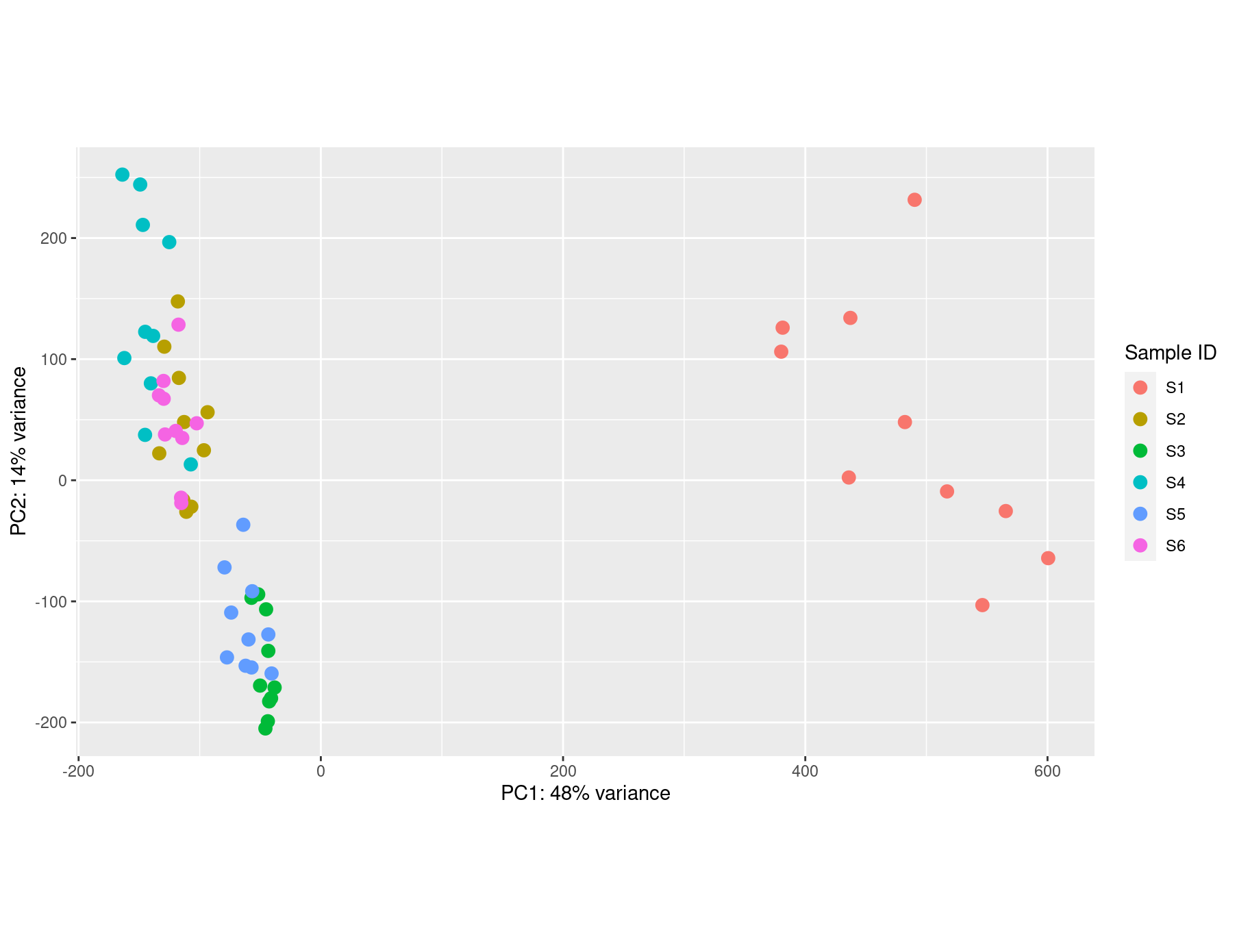
S1 is showing a batch effect compared to all the other samples, and therefore my PCA plot comes out like this.
I wanted to control this batch effect, so I set my design like this.
dds <- DESeqDataSetFromMatrix(countdata,
colData = coldata,
design = ~ S_ID + Condition)
However, I got the error
Error in checkFullRank(modelMatrix) :
the model matrix is not full rank, so the model cannot be fit as specified.
One or more variables or interaction terms in the design formula are linear
combinations of the others and must be removed.
I understand that this error is because my samples are nested within the condition groups. However, even after I read the manual I am not sure how to redesign my data to account for each samples
I appreciate any suggestions or input you can give me. Thank you in advance!