Hello! Hoping I could find some answers here. I have this experimental setup shown in this image:
I was wondering using the formula design = ~ batch + condition would be sufficient enough to control for batch effect considering the fact that the control group is not represented in the other batch. How should I proceed with further analysis? Surprisingly, Limma was able to somewhat correct the data to cluster broadly by condition. I am stuck on what to do.
I will appreciate any answer. Thank you so much!

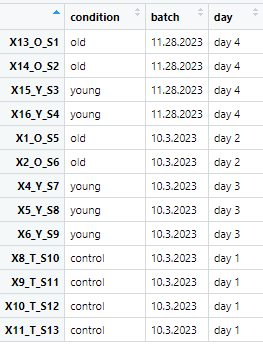

The "batch" is our isolation date for RNA seq, while "day" is the dates we obtained our samples.
Different isolation days will cause a, batch effect. Next time, the experiment needs to be planned more carefully.