Hello,
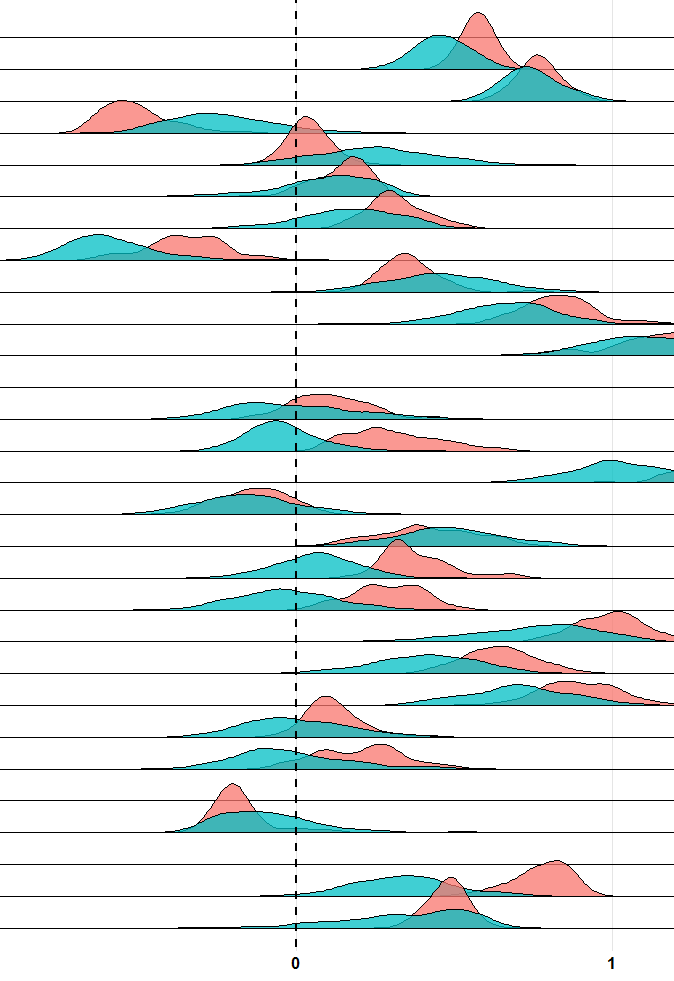
After I plot density of ssGSEA method result, I am curious what the zero line means. As the documentation says,
GSVA assesses the relative enrichment of gene sets across samples using a non-parametric approach.
If the mean of distribution locate at left of zero, does it mean this gene list of interest relatively un-enrich comparing with other genes in the genome???
Best,
Shixiang


Dear Robert,
Thanks for you quick reply. Now I understand what the zero means. So if the mean of enrichment score less much than zero, does it mean that the distribution of the genes inside the gene set rank significantly low than the distribution of genes outside the ranking? Conversely, if the mean of enrichment score greater much than zero, the distribution of the genes inside the gene set rank higher?
Best,
Shixiang
Yes. The meaning of GSVA negative scores has been in fact already asked in this support site in this Does GSVA enrichment score comes negative?.
cheers,
robert.