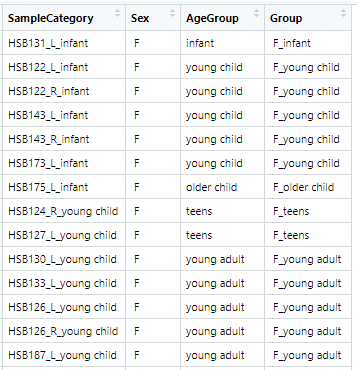
Hi, I am learning how to use limma for bulk RNA-seq data. Below is a snapshot of the metadata file.
My samples are categorised by Sex (Male and Female) and by 6 Age categories (Infant, Young child, Older child, Teens, Young adults and Older adults).
I would like to identify which genes are differentially expressed for each sex and age group. For example, which genes are overexpressed in female infants vs male infants. All the expression values are from controls there is no treatment here.
After reading the limma user guide and several other online resources, the following is the matrix I designed.
Specifying the linear model
design <- model.matrix(~0+ Group, data = metadata)
colnames(design) <- c("F_infant","F_olderadult" ,"F_olderchild", "F_teens","F_youngadult",
"F_youngchild" , "M_infant", "M_olderadult", "M_olderchild","M_teens",
"M_youngadult","M_youngchild")
fit<-lmFit(eset, design)
Contrast Matrix
cont.matrix <- makeContrasts( infants = F_infant-M_infant,
youngchild = F_youngchild - M_youngchild,
olderchild= F_olderchild- M_olderchild,
teens= F_teens-M_teens,
youngadult= F_youngadult - M_youngadult,
olderadult= F_olderadult - M_olderadult,
levels=design)
Fit the model
fit2 <- contrasts.fit(fit, cont.matrix)
fit2 <- eBayes(fit2)
results <- decideTests(fit2, p.value = 0.05)
Can you please let me know if this is right for the design matrix or am I missing something. Thanks in advance for any help!


This seems right.
Thank you for your help! Really appreciate it!