I am after some of the definitions used for KEGG Pathway Edges. In particular — activation, inhibition, expression, repression, and indirect effect. Has anyone come across documentation or a definition guide on how these edges are described and used?
If we are going to use these models beyond graphical use, modeling, or programming purpose, what is the difference between activation vs. expression, and inhibition vs. repression? etc

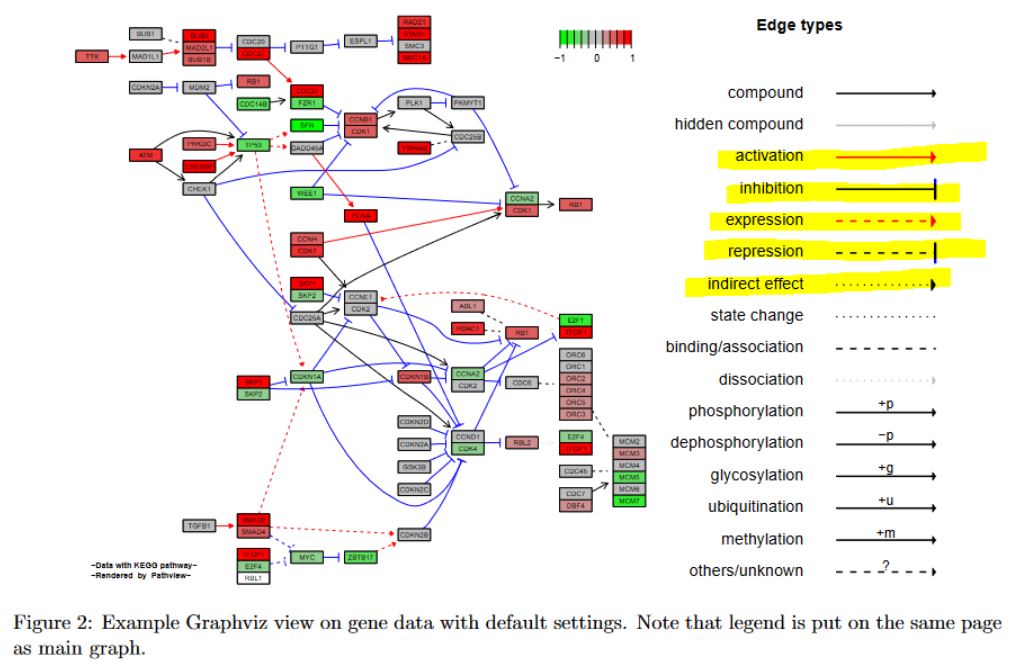

Thanks a heap, deeenes This is helpful :)