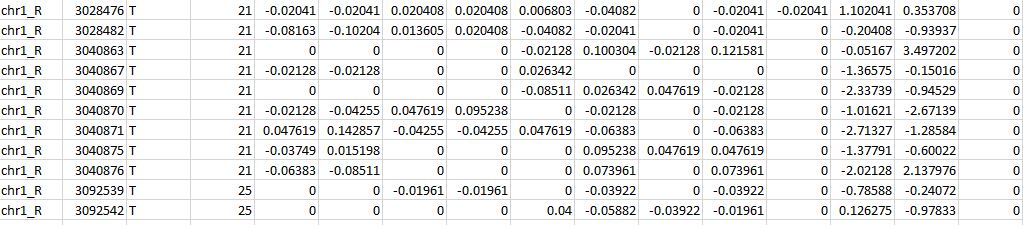
I am trying to find modifications in genomic location. Now, the output I am getting is in the chromosome and location of the modification (attached file). I am not sure how to convert it into a gene name using biomart or any other tool. Would you mind if you please helping me in fixing this issue? Here I don't have a range of base pairs. However, as shown below, I have just a position number in a particular chromosome number.
Convert Chromosome postion to gene symbol
0
Entering edit mode
Similar Posts
Loading Similar Posts
Traffic: 813 users visited in the last hour
Use of this site constitutes acceptance of our User Agreement and Privacy Policy.


What species? What genome build? Do you only want the gene if the variant is within the gene, or close? And if 'close', how far is 'close enough'?
It's human data. Hg19 assembly. I only want the gene if the variant is within the gene.